
 |
New Binary Search for StripesOn Page 1 of this series we described how the MEBES pattern file is organized. We focused more on the geometry that on the database. In order to speed up the program we need to delve deeper into the database organization and make better use of what we learn about the database. 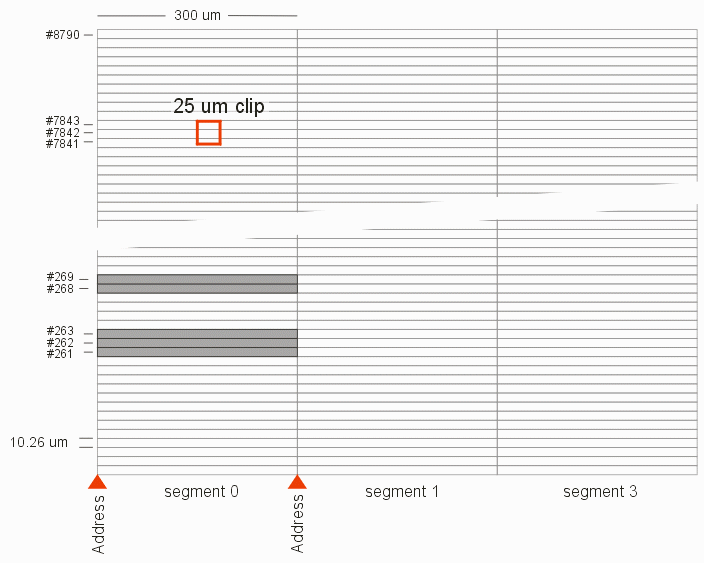 Search LogicGiven: the coordinates of the clip window. We want to extract the trapezoids in this window and either rasterizer them or convert them to GDSII. Our goal is to complete this in 100 milliseconds or less without preloading the 100 GB MEBES file or relying on data transfer faster than 100 MBytes/sec. Read: the job deck transformations and the MEBES header information which includes a segment table with address offsets, the width and height of a stripe. These are both tiny amounts of data and can be read almost instantaneously, even from a remote system. Computation: using the job deck transformations and information in the header of the MEBES data we can quickly (in μsecs) compute which segment (or segments) the clip crosses, as well as which stripes (by index number) the clip crosses. (We know the segment width and we know the stripe height from the MEBES header.) In the example above, we've quickly determined that the clip lies in Segment 0 and that it crosses the stripes with index #7843,#7844 and #7845. Problem: While we can compute the desired stripe index numbers, we don't know where in the stream of bytes that constitute the MEBES pattern file they are located! This is because each stripe contains a different number of bytes. You could load the entire segment 0 into memory to do your search -- but this is a show stopper. A typical segment in a large 50-200 GByte MEBES file will be greater than 20 MB. At 100 MB/sec transfer rate it would take 200 milliseconds to move the data from the file server to the workstation. We've already blown past our timing budget. You could start reading/searching from the beginning of the segment until you find the desired stripe (like we do now) but the farther up in Y the clip is located the more stripes one has to search through. Such an approach can take up to several seconds for clips near the "top" of the layout. Solution: Binary Search
So our program requests a few KB half-way in between those two addresses and it extracts the first stripe index it finds. The stripe index is either too high or too low (unless you are very lucky.) If the index is too high you request a few KB half way above - if too low you request a few bytes half way below. Again mbs2tiff extracts the stripe index and figures out, "Am I above or below the desired stripe index?" Eventually mbs2tiff hits the desired stripe and at that point it is straight forward to read the stripe data from disk, pass the trapezoids to the rasterizer or Boolean engine and produce the requested clip output. Since we're expecting no more than say 50,000 trapezoids in the three stripes we're processing and each trap is about 11 bytes, we're not downloading more than 500 KByte per clip. At 100 MByte/sec transfer rate we are talking about 10-20 msec to get the trapezoid data into the workstation's memory. ConclusionWe think 10 clips per seconds for a 100 GB MEBES file located on a file server connected via a 1 Gb/sec connection to the workstation can be done using the binary search just described. The binary search delays are likely dominated by the disk's seek time as opposed to data transfer rates. Most server disk drives include high performance disks, data striped across multiple disks, large file caches. Since we are only searching on a per segment basis -- so we are not ranging over more than 50-100 MBytes per clip -- the disk cache should be very effective. We already know from our experimental measurements, that for bitmap rasterization, the bottleneck is not the rasterizer. We are checking now whether our Boolean engine and GDSII formatter will run faster than the program can extract trapezoid data. |