Remove Cells (Structures)
A user may want to remove one or more cells from a GDSII file. The command line option, -remove filename, deletes cell definitions listed in the file. If the number of cells to remove is large and they all share some common string, one can also use a regular expression in place of the list.
Command Line Syntax
gdsfilt input_file output_file structure -remove filename
where
input_file gdsii input file
output_file gdsii file to create
structure input file structure to process
-remove filename remove structures with names matching
those in the text file.
Remove File Contents
The rename file has two entries per line. The first entry must match (exactly - case sensitive) the name of a structure in the input file. The second entry will be the new name in the output file.
CELL_1 CELL_A CELL_2 CELL_B
If for some reason (since it is really not supposed to happen) you have structure names with spaces in them then you must use quotes to delimit input and output names.
"CELL 1"CELL A" "CELL 2"CELL B"
Unique Names
There must be unique relationship between input and output names. Two input names must not be mapped to a single output name (even inadvertently). Doing so will completely garble the resulting output file.
Example
As an example we will use our very small file demo1.gds. Below you can see a screen shot of demo 1 along with the structure list (using Qckvu).
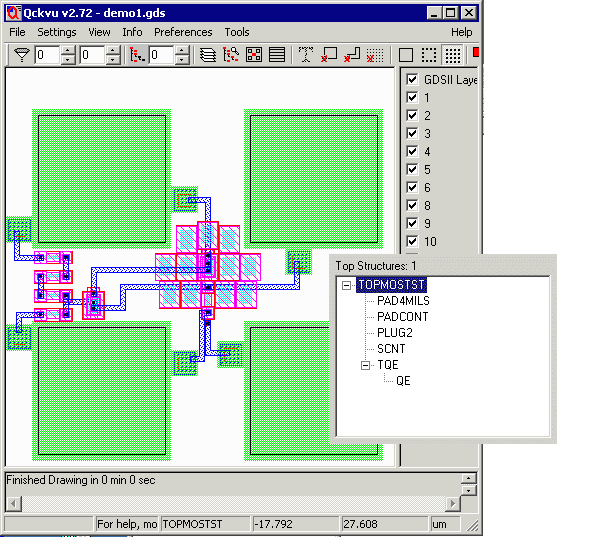
Now we will produce three different output files using the following command line and rename files:
This first example shows how to handle spaces in the structure names.
gdsfilt demo1.gds demo1_ex1.gds = -rename example1.lst
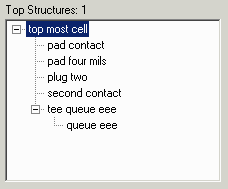
where example1.lst contains: "TOPMOSTST"top most cell" "PAD4MILS"pad four mils" "TQE"tee queue eee" "PADCONT"pad contact" "SCNT"second contact" "PLUG2"plug two" "QE"queue eee" |
|
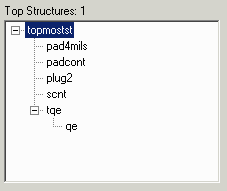
The second example changes the names from upper case to lower case.
gdsfilt demo1.gds demo1_ex2.gds = -rename example2.lst
where example2.lst contains: TOPMOSTST topmostst PAD4MILS pad4mils TQE tqe PADCONT padcont SCNT scnt PLUG2 plug2 QE qe |
|
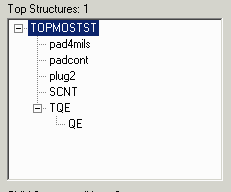
The third example only changes some of the names.
gdsfilt demo1.gds demo1_ex3.gds = -rename example3.lst
where example3.lst contains: PAD4MILS pad4mils PADCONT padcont PLUG2 plug2 |
|